Research Abstract
全ゲノムとエキソームシークエンシングデータからSNVと挿入欠失を検出するための実用的手法
A practical method to detect SNVs and indels from whole genome and exome sequencing data
2013年7月8日 Scientific Reports 3 : 2161 doi: 10.1038/srep02161
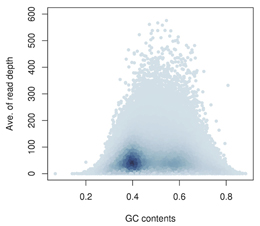
近年の超並列シークエンシング技術の発展により、遺伝的多様性の包括的なカタログの作成が可能になりつつある。しかし、短いリード配列データの比較的高いエラー率のため、高精度で多様性を検出するには洗練された解析手法が必要である。今回、我々は、サンプルごとの全ゲノムシークエンシング(WGS)や全エキソームシークエンシング(WES)データから、一塩基多様性(SNV)および短い挿入や欠失(indel)を検出するための多項確率論手法を開発した。DNAジェノタイピングアレイを用いた評価を行ったところ、コールの一致率がWGSで99.98%、WESで99.99%であることが示された。不一致のコールについてサンガー法によるシークエンシングを行った結果、偽陽性率と偽陰性率が、WGSで0.0068%と0.17%、WESで0.0036%と0.0084%となり、高精度であることが分かった。さらに、短い挿入欠失も高精度で検出できた(WGSで94.7%、WESで97.3%)。我々は、この手法を用いることでヒト疾患がより大きく理解されると確信している。
重水 大智1, 藤本 明洋1, 秋山 真太郎1, 阿部 哲雄1, 中野 かおる2, Keith A. Boroevich1, 山本 裕二郎2, 古田 繭子2, 久保 充明3, 中川 英刀2 & 角田 達彦1
- 理化学研究所 統合生命医科学研究センター 医科学数理研究グループ
- 理化学研究所 統合生命医科学研究センター ゲノムシーケンス解析研究チーム
- 理化学研究所 統合生命医科学研究センター 基盤技術開発研究グループ
The recent development of massively parallel sequencing technology has allowed the creation of comprehensive catalogs of genetic variation. However, due to the relatively high sequencing error rate for short read sequence data, sophisticated analysis methods are required to obtain high-quality variant calls. Here, we developed a probabilistic multinomial method for the detection of single nucleotide variants (SNVs) as well as short insertions and deletions (indels) in whole genome sequencing (WGS) and whole exome sequencing (WES) data for single sample calling. Evaluation with DNA genotyping arrays revealed a concordance rate of 99.98% for WGS calls and 99.99% for WES calls. Sanger sequencing of the discordant calls determined the false positive and false negative rates for the WGS (0.0068% and 0.17%) and WES (0.0036% and 0.0084%) datasets. Furthermore, short indels were identified with high accuracy (WGS: 94.7%, WES: 97.3%). We believe our method can contribute to the greater understanding of human diseases.

