Research Abstract
蛍光標識した生体微小組織のレーザー切り抜き法(LMD法)で明らかとなったマウス海馬領域における遺伝子活性化パターン
Fluorescence laser microdissection reveals a distinct pattern of gene activation in the mouse hippocampal region
2012年11月7日 Scientific Reports 2 : 783 doi: 10.1038/srep00783
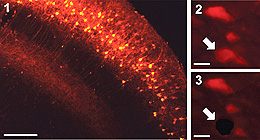
遺伝子発現解析によって組織内の特定の領域や細胞の機能を明らかにする際に、その領域や細胞の組織学的な位置付けを同時に知る必要がある。今回、蛍光標識した組織切片中で標的と定めた領域(ROI)をレーザーで切り抜き、その組織断片中の微量のmRNAの絶対量を定量する方法と組み合わせた画期的な技術革新が実現した。この蛍光LMD定量的RT-PCR法は、最小では単一細胞相当のROIについて、3桁のダイナミックレンジを備えている。海馬の細胞数百個に相当するROIで、最初期遺伝子群の発現レベルを絶対量測定したところ、マウスを飼育ケージから新たな環境に移すと、海馬領域(CA1、CA3、DG)ごとに異なるプロフィルを持つ活性化が起こることがわかった。さらに、CA1領域の遺伝子発現パターンは、各遺伝子の通常の発現量と刺激による遺伝子発現増加率との関係がべき乗則に従っていることがわかった。
吉岡 亘1, 遠藤 のぞみ1, 倉繁 秋江1, 蓜島 旭1*, 遠藤 俊裕1, 柴田 敏幸1, 3, 西山 龍太郎2, 掛山 正心1 & 遠山 千春1
- 東京大学医学系研究科 疾患生命工学センター 健康環境医工学部門
- ライカマイクロシステムズ株式会社 リサーチ・クリニカル事業部
- 東京大学 大学院医学系研究科 国際保健学専攻 人類生態学教室
* 現所属: 群馬大宅医学系研究科 応用生理学分野
A histoanatomical context is imperative in an analysis of gene expression in a cell in a tissue to elucidate physiological function of the cell. In this study, we made technical advances in fluorescence laser microdissection (LMD) in combination with the absolute quantification of small amounts of mRNAs from a region of interest (ROI) in fluorescence-labeled tissue sections. We demonstrate that our fluorescence LMD-RTqPCR method has three orders of dynamic range, with the lower limit of ROI-size corresponding to a single cell. The absolute quantification of the expression levels of the immediate early genes in an ROI equivalent to a few hundred neurons in the hippocampus revealed that mice transferred from their home cage to a novel environment have distinct activation profiles in the hippocampal regions (CA1, CA3, and DG) and that the gene expression pattern in CA1, but not in the other regions, follows a power law distribution.

