The Missing Kingdom: Why Fungi Must Be Central to Conservation Strategy
28 December 2025
Published online 7 January 2019
Genomic analyses reveal cholera strain’s weak points and likely entry paths to the country.
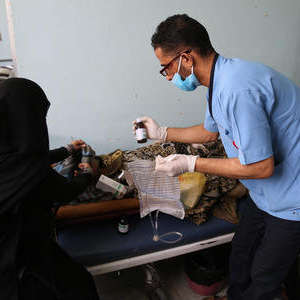
Lack of access to safe water and basic sanitation in war-torn Yemen has resulted in more than one million suspected cholera cases with almost 2,500 deaths from the disease.
To understand the nature of this epidemic, researchers at France's Institut Pasteur and Britain's Wellcome Sanger Institute, together with collaborators from Egypt, Iran, Iraq, Saudi Arabia, the United Arab Emirates and Yemen, sequenced the genome of 42 Vibrio cholerae infected samples, collected from patients in Yemen and nearby regions. These were compared to 74 other samples from South Asia, the Middle East, Eastern and Central Africa, and a global collection of over 1,000 samples.
The results showed that the culprit of the 2016 and 2017 outbreaks in Yemen belongs to the seventh cholera pandemic that began in 1961 in Indonesia and spread globally. Within this bacterial lineage, this specific strain of V. cholerae is related to recent outbreaks in Eastern Africa, but different from those that have been circulating in the Middle East over the last decade, suggesting its route to Yemen via Africa.
“The International Organization for Migration reported that, every year, thousands of migrants from East Africa travel to Yemen looking to reach Gulf countries. Obtaining genetic samples from neighbouring Somalia and Ethiopia, where two large cholera outbreaks occurred in 2016, would be key to precisely reconstruct the route of transmission of this strain,” says mathematical modelling epidemiologist Anton Camacho of Epicentre in France, who was not involved in the study.
The team also discovered that the Yemeni cholera strain is more susceptible to antibiotics than other strains of the seventh cholera pandemic. “To see a V. cholerae strain that had lost many resistance genes to become only resistant to a few old generation antibiotics was very unusual,” says the study’s corresponding co-author, François-Xavier Weill of the Institut Pasteur in France.
In particular, unlike other strains of the same lineage, a small genetic variation makes this strain susceptible to the antibiotic polymyxin. “V. cholerae strains are undergoing constant cryptic changes in the genome, which may influence virulence, transmission and spread,” comments Asish Mukhopadhyay of ICMR-National Institute of Cholera and Enteric Diseases in India, who was not involved in the study.
“These findings underscore the importance of coordinated global cholera surveillance efforts to understand, track and prevent the spread of cholera through populations,” adds epidemiologist Molly Franke of Harvard Medical School, USA, who was also not involved in the study.
doi:10.1038/nmiddleeast.2019.2
Weill, F-X., et al. Genomic insights into the 2016–2017 cholera epidemic in Yemen. Nature https://doi.org/10.1038/s41586-018-0818-3 (2019).
Stay connected: