The Missing Kingdom: Why Fungi Must Be Central to Conservation Strategy
28 December 2025
Published online 21 November 2016
Gene therapy partially restores vision in rats. The new technique enables scientists to repair genes in adult tissues.
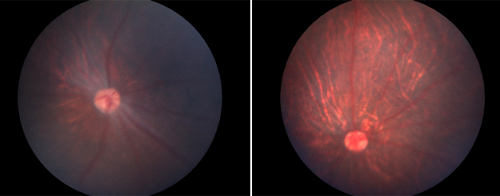
© Keiichiro Suzuki
Their approach holds great promise for treating inherited diseases.
To repair the rats’ genetic defect, the team, which included researchers from Saudi Arabia’s King Abdullah University of Science and Technology (KAUST), developed a new method which allowed them to introduce DNA at a targeted location in the non-dividing cells which make up most adult tissues and organs, such as neurons and muscles.
The new approach builds on the genome-editing technique CRISPR-Cas9, which co-opts an immune mechanism from bacteria to enable scientists to direct a payload to a specified target location in the genome. In tandem with native DNA repair mechanisms, this has brought precise genome-editing within scientists’ reach.
So far these technologies have primarily been used to knock out genes rather than insert DNA at a specific location. For example, in a study published this year, CRISPR-Cas9 was used to disable one of the malfunctioning genes in rats with the inherited degenerative eye disease, retinitis pigmentosa, preventing degradation of the rats’ retinas.
A major shortcoming has been the restriction of these techniques to dividing cells. To overcome these limitations, the team developed a strategy called homology-independent targeted integration (HITI) which combines a customized nucleic acid cocktail with CRISPR-Cas9 to take advantage of a DNA repair mechanism that is active even in non-dividing cells. They tested the new technique by using it to introduce DNA encoding a green fluorescent protein (GFP) downstream of a specific gene in cultured human neurons.
“To our excitement, a couple of days later, we found many neurons with GFP! That was the first indication that HITI could work in non-dividing cells,” says Juan Carlos Izpisua Belmonte of the Salk Institute in California, who led the project. “We immediately designed a series of experiments and applied HITI directly in vivo using rodent models.”
The next test was to use HITI to introduce the GFP sequence into muscle, kidney and brain cells in live mice.
The team used different techniques to deliver the HITI cocktail to these tissues, including local and systemic injections of a viral vector called AAV carrying the HITI components. All of the experiments worked, though the efficiency varied depending on the tissue and delivery mode. “The development of more efficient gene delivery methods, which is a bottleneck for in vivo genome editing in general, will improve HITI efficiency in the near future” says Izpisua Belmonte.
As a final test, the researchers used HITI to repair the defective Mertk gene in the eyes of three-week-old rats with retinitis pigmentosa, which causes blindness. “We introduced a functional copy of the gene at the right genome location, thereby repairing the disease-causing mutation,” says Izpisua Belmonte.
Five weeks later, the rats were able to detect light, and tests showed that they had significantly recovered their visual functions.
“This is a very compelling paper,” says Richard Jude Samulski, director of the Gene Therapy Center at the University of North Carolina at Chapel Hill, who wasn’t involved in the study. “The new approach accomplishes what people wanted to do in the early days of genetic engineering, which was to swap out a mutation for healthy DNA. It sets us in a direction which I think many people will follow.”
The ability to correct loss-of-function mutations in adult tissues holds obvious therapeutic appeal, and HITI also has the potential to advance basic and translational research. However, the technique still needs refinement, and Izpisua Belmonte also notes that its safety needs to be studied. “HITI has both error-prone and error-free activities, and further understanding the detailed molecular mechanisms underlying each will help us design a more effective strategy to maximize its error-free activity for applications in humans.”
doi:10.1038/nmiddleeast.2016.207
Stay connected: