The Missing Kingdom: Why Fungi Must Be Central to Conservation Strategy
28 December 2025
Published online 21 February 2019
A reliable, reproducible way to modify the roundworm’s genome promises to be a game changer for genetic studies.
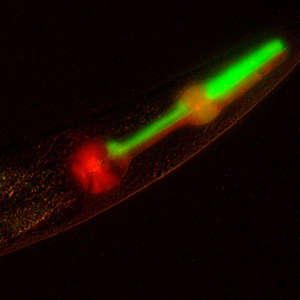
© 2018 Karen Lynn Artiles
Now, researchers at Stanford University Medical School and King Abdullah University of Science and Technology (KAUST) have demonstrated1 a way to engineer the worm’s genome so scientists can probe even more deeply into life at its most fundamental level. They report a strategy to create new populations of C. elegans that do not follow the rules of classical, or Mendelian, genetics.
In contrast to offspring that inherit DNA from both parents, the new populations only inherit DNA from the paternal line.
This non-Mendelian inheritance pattern is of major interest to scientists exploring gene function, evolutionary biology and epigenetics—the study of how genes are turned on or off without changes in DNA sequence.
The researchers designed the worms with the desired inheritance pattern efficiently, in a single generation, and reproducibly, with a high (80 per cent) success rate.
“The surprising ability to re-engineer the fundamental Mendelian inheritance pattern raises exciting possibilities,” says Christian Frøkjær-Jensen, a co-author at KAUST, who was a visiting scientist at the Stanford University laboratory headed by Nobel laureate Andrew Fire, a collaborator on the study.
“For example, the ability to decouple genomic DNA inheritance from the inheritance of everything else in the fertilized egg, such as mitochondria and RNA, will enable us to understand how epigenetic patterns are initiated and maintained,” he says.
The study builds on a breakthrough2 by Judith Besseling and Henrik Bringmann at the Max Planck Institute for Biophysical Chemistry in 2016. By engineering a gene called GPR1, they found a way of exerting pulling forces during the first division of C. elegans to generate worms with non-Mendelian inheritance at the two-cell stage. However, the engineered gene was prone to being turned off, or silenced, making it difficult to achieve consistent results. Visual confirmation of the inheritance pattern also proved challenging.
To overcome these issues, the new method incorporated other DNA segments to counteract the silencing effect. The resultant worms’ GPR1 gene was not only resistant to silencing, but the paternal inheritance pattern was also easy to confirm at low magnification.
Molecular cell biologist Arshad Desai of the University of California, San Diego, who was not involved in the study, says the new method represents “a significant technical effort to improve the ability to manipulate forces in the first division and thereby produce uniparental genome-derived individual cells in two-cell-stage embryos.”
The researchers expect their method could become a routine tool for the C. elegans research community and a resource for scientists working on other model systems.
“The wider implications for basic research are that cherished, fundamental laws, such as Mendel's inheritance laws, are not as immutable as one might expect,” Frøkjær-Jensen says.
doi:10.1038/nmiddleeast.2019.26
Stay connected: