The Missing Kingdom: Why Fungi Must Be Central to Conservation Strategy
28 December 2025
Published online 19 July 2016
Researchers find a bias in the process of elimination that causes some cells to form antibody sites while others are cast off.
© Mohamed Khass and Harry Schroeder Jr.
This new understanding of the regulation of B-cell development has shown that the process of creating the antibody repertoire is biased and selects for specific categories of antigen recognition structures.
Researchers in the United States, Egypt and Germany have discovered that one of the earliest selective checkpoints in B-cell development participates in an elimination process. This process is based on the presence of certain amino acid chains at specific positions within the antigen binding site of the immunoglobulin.
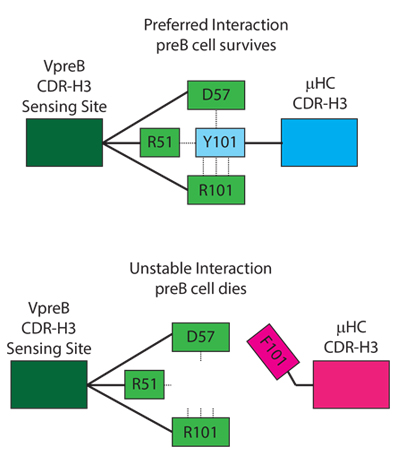
Using mice, they examined B cells that were being tested by this early checkpoint and found that cells with antigen binding sites that are rich in tyrosine and present at specific positions were allowed to live, whereas those expressing hydrophobic amino acids were being killed.
This early checkpoint process has positive and negative outcomes. While an earlier screening means B cells can produce antibodies that can fight common pathogens more efficiently, it also means that the elimination process rejects cells that can be needed for more complicated viruses.
“Some of the antibodies that protect against HIV have features that would normally be selected against at this early selection step. We now have a new explanation for why the immune system responds easily to some microbes and has more difficulty responding to others,” says Harry Schroeder, co-author of the paper.
Future studies can address how these selection stages can be manipulated within the cell development, adds Schroeder.
doi:10.1038/nmiddleeast.2016.109
Stay connected: