The Missing Kingdom: Why Fungi Must Be Central to Conservation Strategy
28 December 2025
Published online 28 July 2011

Hardy bacteria adapted to the extreme conditions found at the bottom of the Red Sea might become a plentiful well of enzymes with numerous applications in biotechnologies.
André Antunes, Uli Stingl and their research team at the Marine Microbial Ecology group at the Red Sea Research Center in the King Abdullah University of Science and Technology (KAUST), Thuwal, Saudi Arabia, set out to explore these species. They have so far isolated and studied several different strains of extremophilic microbes, which are microbes that live in harsh conditions of high temperature and salinity, from an underwater brine pool lying 1.4 km below the surface of the Red Sea.
These brine pools are among the most extreme and inhospitable environments on Earth with high pressures, temperatures that range from 50 – 70 °C, little to no light or oxygen, and high concentrations of salt and heavy metals.
The Rea Sea brine pools remain largely unexplored and, until recently, were thought to be devoid of life. They are, in fact, teeming with microbes, some of which represent entirely new bacterial lineages, according to the researchers' phylogenetic studies.
Shaban Deep is one of 25 deep brine pools in the Red Sea's central basin. "[It]is a remarkable environment with very exciting microbiology," says Mike Dyall-Smith who studies Halobacteriaceae at Charles Sturt University in New South Wales, Australia. "Its biology is likely to be very different from other known environments."
Antunes and his colleagues isolated the bacteria from samples of sea floor sediment collected during an expedition of the German oceanographic research vessel RV Meteor, which holds 20 well-equipped laboratories.
"Some of the most interesting microbes from Shaban Deep are Halorhabdus tiamatea, Haloplasma contractile and Salinisphaera shabanensis," says Antunes. " H. tiamatea is the first member of the Archaea ever isolated from these environments, while S. shabanensis and H. contractile are the first members of two new bacterial orders."
Halorhabdus tiamatea is related to the archaea Halorhabdus utahensis found in the Great Salt Lake in Utah, United States, but shows specific adaptations to life in the deep sea brines. It lacks pigmentation and has a preference for growing in the absence of oxygen.
Salinisphaera shabanensis lives at the interface between brine and seawater, where drastic changes in environmental conditions and nutrient availability are observed. Salinity in seawater is around 3%, but it can be as high as 27% in the brine pools. Accordingly it has a flexible physiology enabling it to cope with life under highly variable conditions.
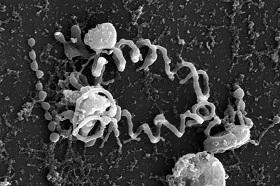
The third microbe the team found in the Red Sea, Haloplasma contractile, forms tentacle-like protrusions that alternate between a straight, relaxed shape and a contracted, corkscrew-like one. Their function is unclear, but Antunes thinks they might cling to surfaces and produce water movements that replenish nutrients in the microenvironment around the cells.
In collaboration with researchers from the American University in Cairo, Antunes and his colleagues have recently sequenced the H. contractile genome. The results, published recently in the Journal of Bacteriology, show that the microbe contains no less than seven copies of genes encoding seven copies of a specific set of genes encoding cytoskeletal proteins.
Until recently, such genes were widely thought to be absent in bacteria, and this is the highest copy number reported in a bacterium to date. Antunes believes they might play an important role in the cell structure and in cellular motility and contraction of the protrusions, which would make H. contractile unique among bacteria.
"I do not know of [bacterial] cells with arms like those of H. contractile," says Dyall-Smith. "When you find biological novelty like this, there is a much greater chance of finding commercially useful products."
Enzymes from extremophilic bacteria are applied widely in the food, pharmaceutical and textiles industries and in research such as holography and computer science. The salt-loving microbes that thrive in the Red Sea's deep brine pools have developed novel metabolic processes, and probably synthesize enzymes that can lend themselves to numerous applications.
"The potential is enormous," says Antunes, "because each brine pool has a unique combination of environmental conditions that would be reflected in very different and unique microbial communities. Most of them have never been studied, so this is just a first glimpse of the enormous biodiversity present there. Some of the most promising applications include production of bioplastics and other biopolymers, and bioremediation for heavy metal and oil contamination."
The researchers are now analyzing the genomes of the mıcrobes they have dıscovered in greater detail to look for interesting enzymes and proteins.
doi:10.1038/nmiddleeast.2011.96
Stay connected: