River subbasins more depleted than
official figures show
09 April 2024
Published online 30 December 2016
Egyptian scientists develop affordable diagnostic technology of gold nanoparticles which uses HCV’s genomic signature.
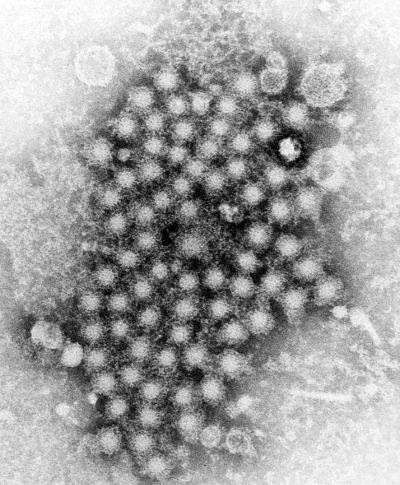
© CDC/ E.H. Cook, Jr.
The technology, the researchers say, is simple, cost-effective and highly sensitive.
In Egypt, 500,000 new cases of Hepatitis C emerge each year — a huge challenge in a country where 15% of the population is already infected with the virus. Existing technologies that can detect and quantify HCV in clinical samples are expensive and remain inaccessible to impoverished populations.
Nanobiotechnology has potential diagnostic applications for a variety of pathogenic viruses and bacteria.
“Despite rapid developments in therapies, diagnosis remains the bottleneck in the fight against HCV,” says Sherif El-Khamisy, the scientist responsible for the new diagnostic tool, which relies on nanomaterials instead of chemicals.
The underlying principle of the technology, according to El-Khamisy, is to couple an HCV-specific probe with gold nanoparticles and then change the behaviour of the particles by influencing their aggregation and inducing a colour change based on the presence or lack of HCV in a clinical sample. A positive sample will typically produce a colour change that can be visualized or quantified.
Other methods typically use salt for inducing the aggregation of gold nanoparticles, but El-Khamisy and colleagues are replacing pH values with gold nanoparticles to induce aggregation in other gold nanoparticles with the opposite charge.
“Current methods such as quantitative PCR are expensive, time-consuming and require well-equipped labs,” explains El-Khamisy. His team’s method is fast and cheap, he says, yielding results in just 30 minutes. “It will hopefully help us reaching millions of people living in rural areas who never had access to excising expensive technologies. The method is almost as sensitive as the gold standard real-time quantititative PCR at about a tenth of its cost.”
El-Khamisy even says that the sensor has some potential as a biomarker detector for use in relation with cancer and several cardiovascular diseases.
The advantage of this technology over similar ones, says Gheyath K. Nasrallah, assistant professor of Biomedical science at the College of Health Sciences in Qatar University, is that it has improved the stability or aggregations of the nanoprobes.
Nasrallah, who was not involved in the study, says that aggregation of nanoparticles relying on pH strength usually produces many false positives – but this technology relies on RNA target folding instead.
He adds that even at 93%, its colorimetric sensitivity “still needs to be improved, especially when it comes for diagnosis of life-threating and highly contagious virus such as HCV”.
Epidemiologist F. DeWolfe Miller, of the John A. Burns School of Medicine University of Hawaii, agrees. Miller believes the new technology can reduce prognosis time and cost, but lacks sensitivity.
Laith J. Abu-Raddad, director of the Biostatistics, Epidemiology, and Biomathematics Research Core at Weill-Cornell Medicine Qatar, says the probe is significant for its public health potential. Nationally, “the availability of assays that can diagnose infection rapidly and at low cost will be a great asset for [national] screening programs.”
El-Khamisy says he will work on validating the nano-sensor for large scale production. “We need to invest in the quality control and quality assurance of the production process as we move forward,” he says.
doi:10.1038/nmiddleeast.2016.225
Stay connected: